Code
library(here)
df_SEACAR <- readr::read_delim(
here("data/Discrete WQ - 10006.txt"),
delim = "|"
)
df_OLD <- readr::read_delim(here::here("data/allDataSEACAR.csv"), delim=",")
library(dplyr)Click on any element for more details.
# A tibble: 26 × 3
value in_SEACAR in_OLD
<chr> <lgl> <lgl>
1 Salinity TRUE TRUE
2 Water Temperature TRUE TRUE
3 pH TRUE TRUE
4 Total Suspended Solids TRUE TRUE
5 Chlorophyll a, Corrected for Pheophytin TRUE TRUE
6 Light Extinction Coefficient TRUE FALSE
7 Colored Dissolved Organic Matter TRUE FALSE
8 NO2+3, Filtered TRUE TRUE
9 Ammonium (NH4) TRUE FALSE
10 Phosphate, Filtered (PO4) TRUE TRUE
11 Dissolved Oxygen TRUE TRUE
12 Dissolved Oxygen Saturation TRUE TRUE
13 Turbidity TRUE TRUE
14 Chlorophyll a, Uncorrected for Pheophytin TRUE TRUE
15 Total Nitrogen TRUE TRUE
16 Total Phosphorus TRUE TRUE
17 Nitrate (NO3) TRUE TRUE
18 Nitrite (NO2) TRUE TRUE
19 Nitrogen, organic TRUE TRUE
20 Total Ammonia (N) TRUE FALSE
21 Nitrogen, inorganic TRUE FALSE
22 Specific Conductivity TRUE TRUE
23 Total Kjeldahl Nitrogen TRUE TRUE
24 Secchi Depth TRUE FALSE
25 Ammonium, Filtered (NH4) FALSE TRUE
26 Ammonia, Un-ionized (NH3) FALSE TRUE library(ggplot2)
library(tidyr)
# Count rows by ParameterName for SEACAR dataset
seacar_parameter_counts <- df_SEACAR %>%
group_by(ParameterName) %>%
count(name = "SEACAR_Count") %>%
arrange(desc(SEACAR_Count))
# Count rows by ParameterName for OLD dataset
old_parameter_counts <- df_OLD %>%
group_by(ParameterName) %>%
count(name = "OLD_Count") %>%
arrange(desc(OLD_Count))
# Create combined dataframe for plotting
combined_parameter_counts <- full_join(
seacar_parameter_counts,
old_parameter_counts,
by = "ParameterName"
) %>%
mutate(
SEACAR_Count = replace_na(SEACAR_Count, 0),
OLD_Count = replace_na(OLD_Count, 0)
) %>%
# Reshape to long format for dodged bars
pivot_longer(
cols = c(SEACAR_Count, OLD_Count),
names_to = "source",
values_to = "count"
) %>%
mutate(source = ifelse(source == "SEACAR_Count", "SEACAR_STD", "OLD"))
# === Split the plot into two because there are many parameters
TEXT_ANGLE = 90
# Split parameters alphabetically
parameters <- unique(combined_parameter_counts$ParameterName)
mid_point <- ceiling(length(parameters) / 2)
first_half <- sort(parameters)[1:mid_point]
second_half <- sort(parameters)[(mid_point + 1):length(parameters)]
# Create first plot (A-M)
plot1_data <- combined_parameter_counts %>%
filter(ParameterName %in% first_half) %>%
mutate(ParameterName = factor(ParameterName, levels = sort(first_half)))
plot1 <- ggplot(plot1_data, aes(x = ParameterName, y = count, fill = source)) +
geom_bar(stat = "identity", position = "dodge") +
labs(
title = "Row Count by ParameterName (A-M)",
x = "ParameterName",
y = "Number of Rows",
fill = "Dataset"
) +
theme_minimal() +
theme(axis.text.x = element_text(angle = TEXT_ANGLE, hjust = 1))
# Create second plot (N-Z)
plot2_data <- combined_parameter_counts %>%
filter(ParameterName %in% second_half) %>%
mutate(ParameterName = factor(ParameterName, levels = sort(second_half)))
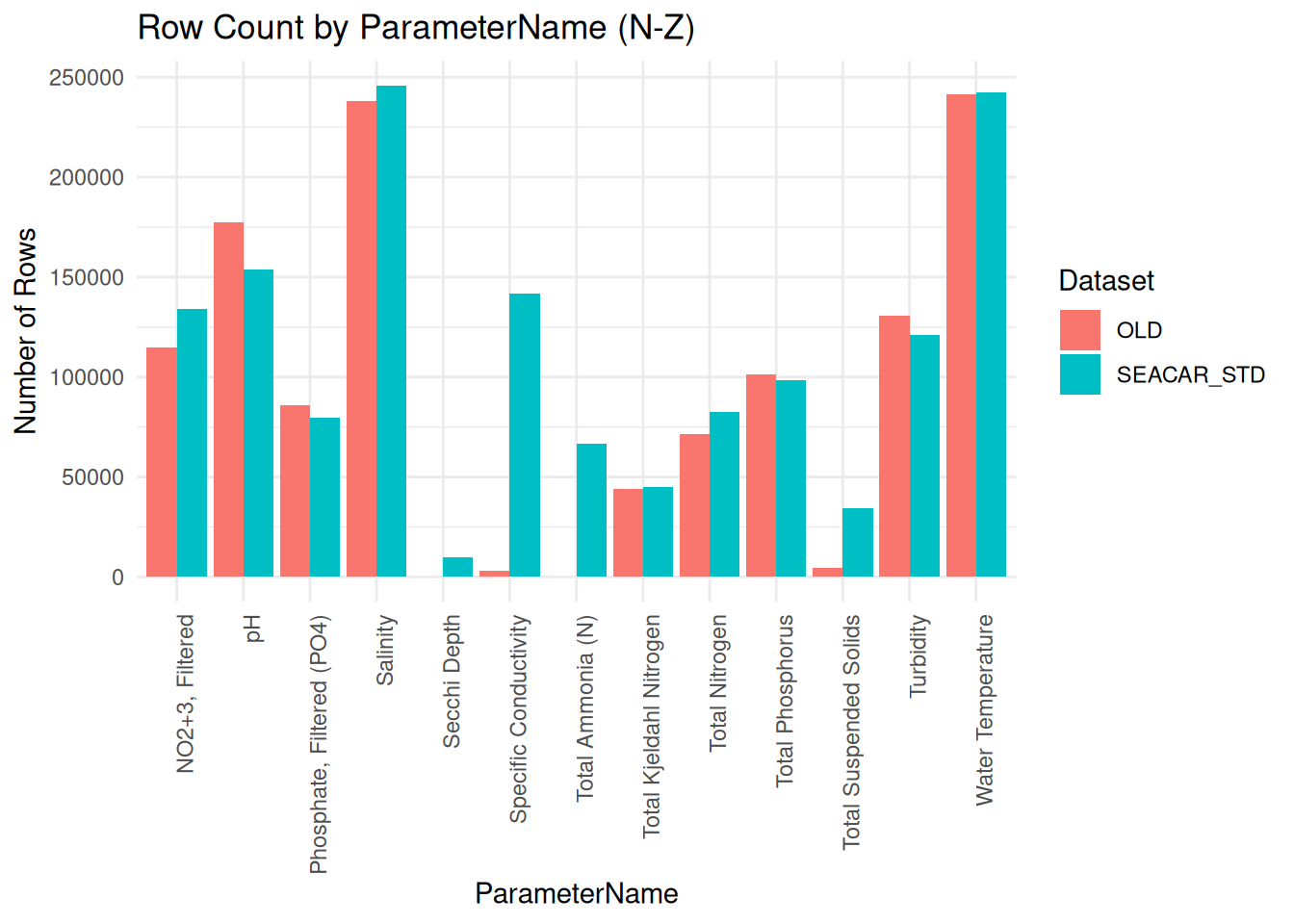
plot2 <- ggplot(plot2_data, aes(x = ParameterName, y = count, fill = source)) +
geom_bar(stat = "identity", position = "dodge") +
labs(
title = "Row Count by ParameterName (N-Z)",
x = "ParameterName",
y = "Number of Rows",
fill = "Dataset"
) +
theme_minimal() +
theme(axis.text.x = element_text(angle = TEXT_ANGLE, hjust = 1))
# Display both plots
print(plot1)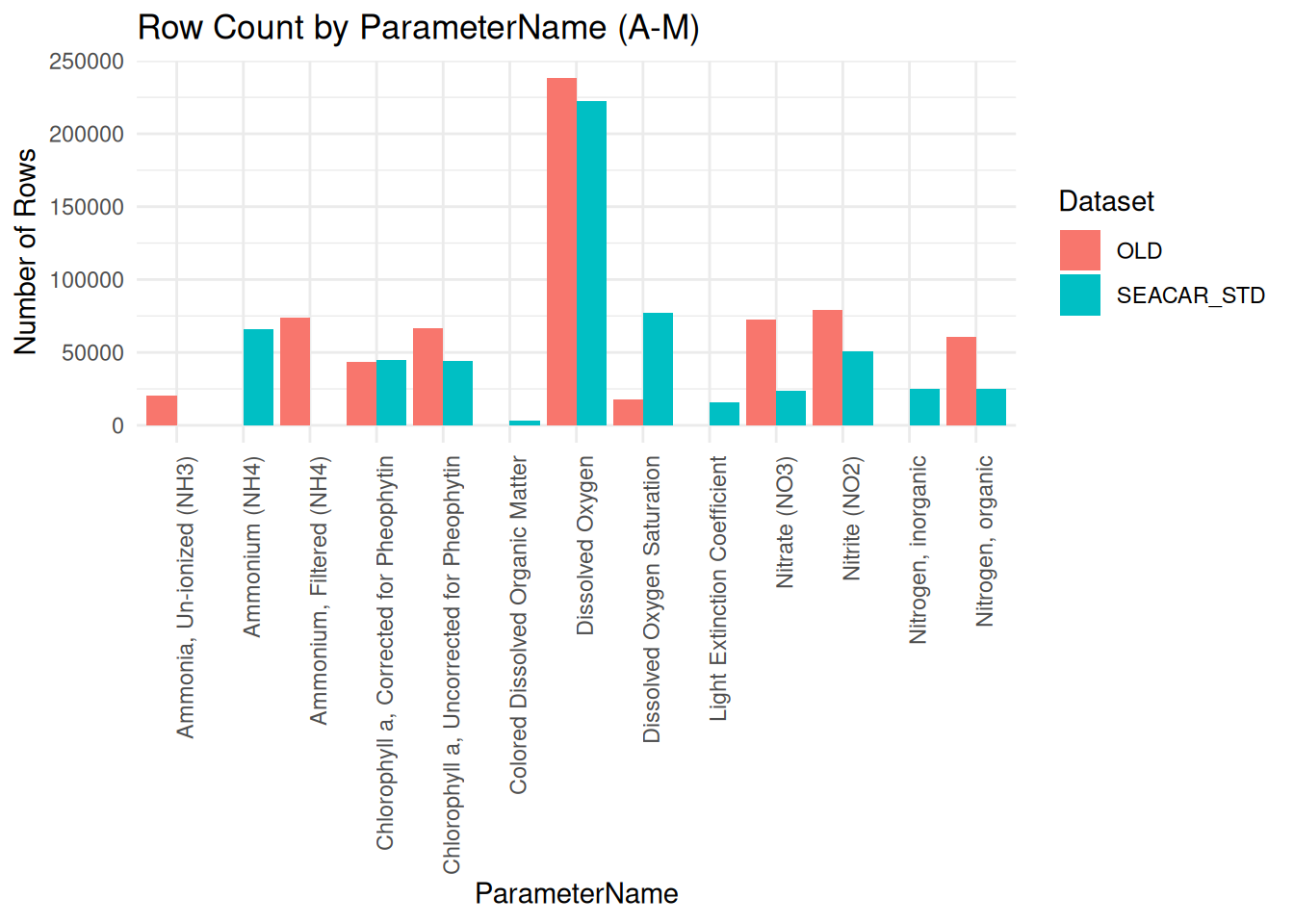
column in_SEACAR in_OLD
1 RowID TRUE TRUE
2 ProgramID TRUE TRUE
3 ProgramName TRUE TRUE
4 Habitat TRUE TRUE
5 IndicatorID TRUE TRUE
6 IndicatorName TRUE TRUE
7 ParameterID TRUE TRUE
8 ParameterName TRUE TRUE
9 ParameterUnits TRUE TRUE
10 ResultValue TRUE TRUE
11 SampleDate TRUE TRUE
12 Year TRUE TRUE
13 Month TRUE TRUE
14 SEACAR_QAQCFlagCode TRUE TRUE
15 SEACAR_QAQC_Description TRUE TRUE
16 Include TRUE TRUE
17 LocationID TRUE FALSE
18 ProgramLocationID TRUE TRUE
19 ProgramLocationName TRUE FALSE
20 OriginalLatitude TRUE TRUE
21 OriginalLongitude TRUE TRUE
22 AreaID TRUE TRUE
23 ManagedAreaName TRUE TRUE
24 AreaID_Buff TRUE FALSE
25 ManagedAreaName_Buff TRUE FALSE
26 EstuarineMarine TRUE FALSE
27 OIMMP TRUE FALSE
28 CHIMMP TRUE FALSE
29 CoralRegion TRUE FALSE
30 ActivityType TRUE TRUE
31 ActivityDepth_m TRUE TRUE
32 RelativeDepth TRUE TRUE
33 TotalDepth_m TRUE TRUE
34 MDL TRUE TRUE
35 PQL TRUE TRUE
36 DetectionUnit TRUE TRUE
37 ValueQualifier TRUE TRUE
38 ValueQualifierSource TRUE TRUE
39 SampleFraction TRUE FALSE
40 ResultComments TRUE TRUE
41 SEACAR_EventID TRUE TRUE
42 ExportVersion TRUE TRUE
43 ...1 FALSE TRUE
44 MADup FALSE TRUE
45 Region FALSE TRUElibrary(ggplot2)
library(tidyr)
library(lubridate)
# Count rows per date for each dataset, bin by year
seacar_counts <- df_SEACAR %>%
group_by(SampleDate = floor_date(SampleDate, "year")) %>%
count(name = "SEACAR_Count") %>%
ungroup()
old_counts <- df_OLD %>%
group_by(SampleDate = floor_date(SampleDate, "year")) %>%
count(name = "OLD_Count") %>%
ungroup()
# Combine and plot
combined_counts <- full_join(seacar_counts, old_counts, by = "SampleDate") %>%
arrange(SampleDate) %>%
mutate(
SEACAR_Count = replace_na(SEACAR_Count, 0),
OLD_Count = replace_na(OLD_Count, 0)
)
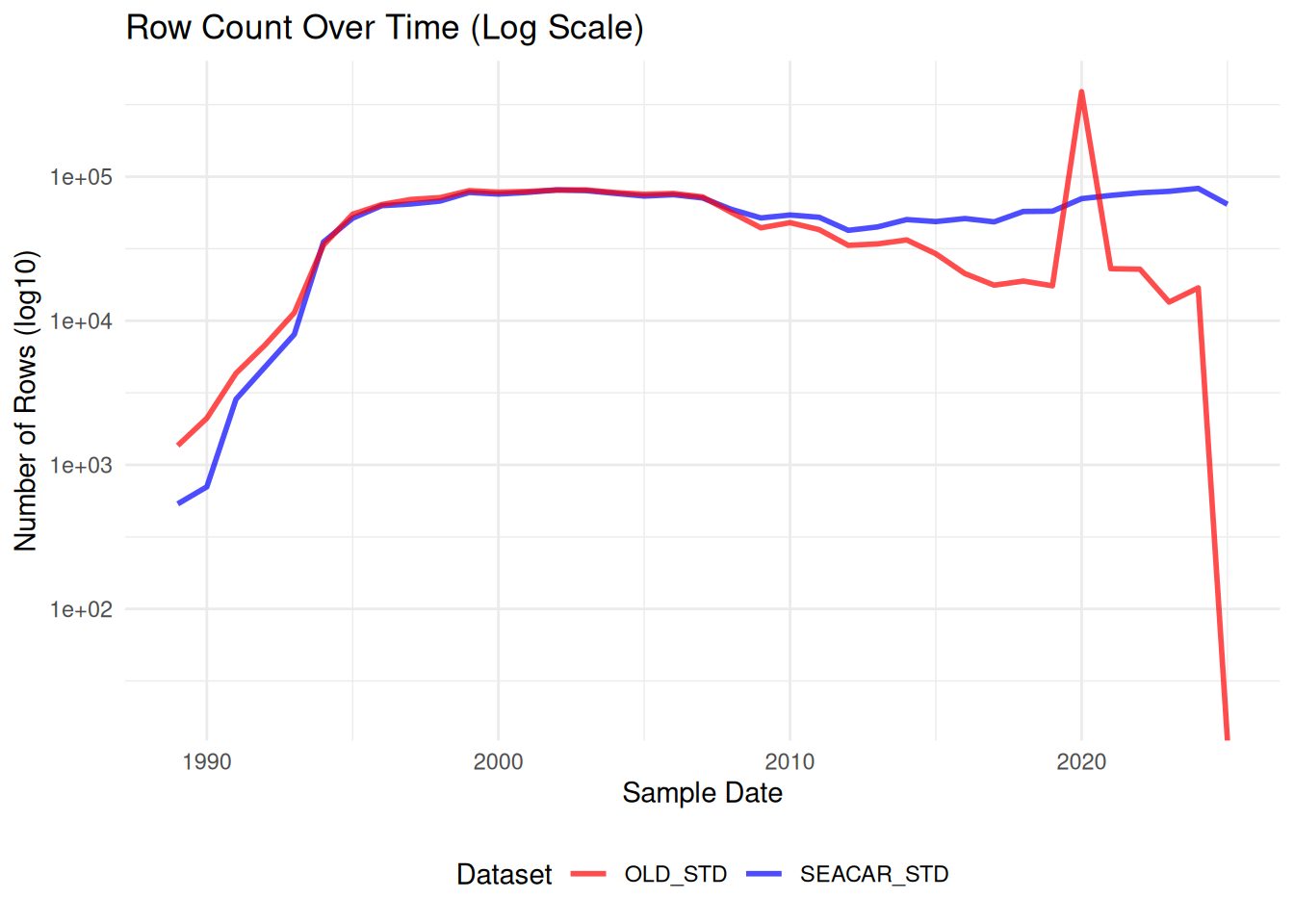
ggplot(combined_counts, aes(x = SampleDate)) +
geom_line(aes(y = SEACAR_Count, color = "SEACAR_STD", linetype = "SEACAR_STD"), size = 1, alpha = 0.7) +
geom_line(aes(y = OLD_Count, color = "OLD_STD", linetype = "OLD_STD"), size = 1, alpha = 0.7) +
scale_y_log10() +
scale_linetype_manual(values = c("SEACAR_STD" = "solid", "OLD_STD" = "solid")) +
scale_color_manual(values = c("SEACAR_STD" = "blue", "OLD_STD" = "red")) +
labs(
title = "Row Count Over Time (Log Scale)",
x = "Sample Date",
y = "Number of Rows (log10)",
color = "Dataset",
linetype = "Dataset"
) +
theme_minimal() +
theme(legend.position = "bottom")