library(ggplot2)
library(tidyr)
library(lubridate)
# Count rows per date for each dataset, bin by year
seacar_counts <- dfs$SEACAR_STD %>%
group_by(SampleDate = floor_date(SampleDate, "year")) %>%
count(name = "SEACAR_Count") %>%
ungroup()
old_counts <- dfs$OLD %>%
group_by(SampleDate = floor_date(SampleDate, "year")) %>%
count(name = "OLD_Count") %>%
ungroup()
# Combine and plot
combined_counts <- full_join(seacar_counts, old_counts, by = "SampleDate") %>%
arrange(SampleDate) %>%
mutate(
SEACAR_Count = replace_na(SEACAR_Count, 0),
OLD_Count = replace_na(OLD_Count, 0)
)
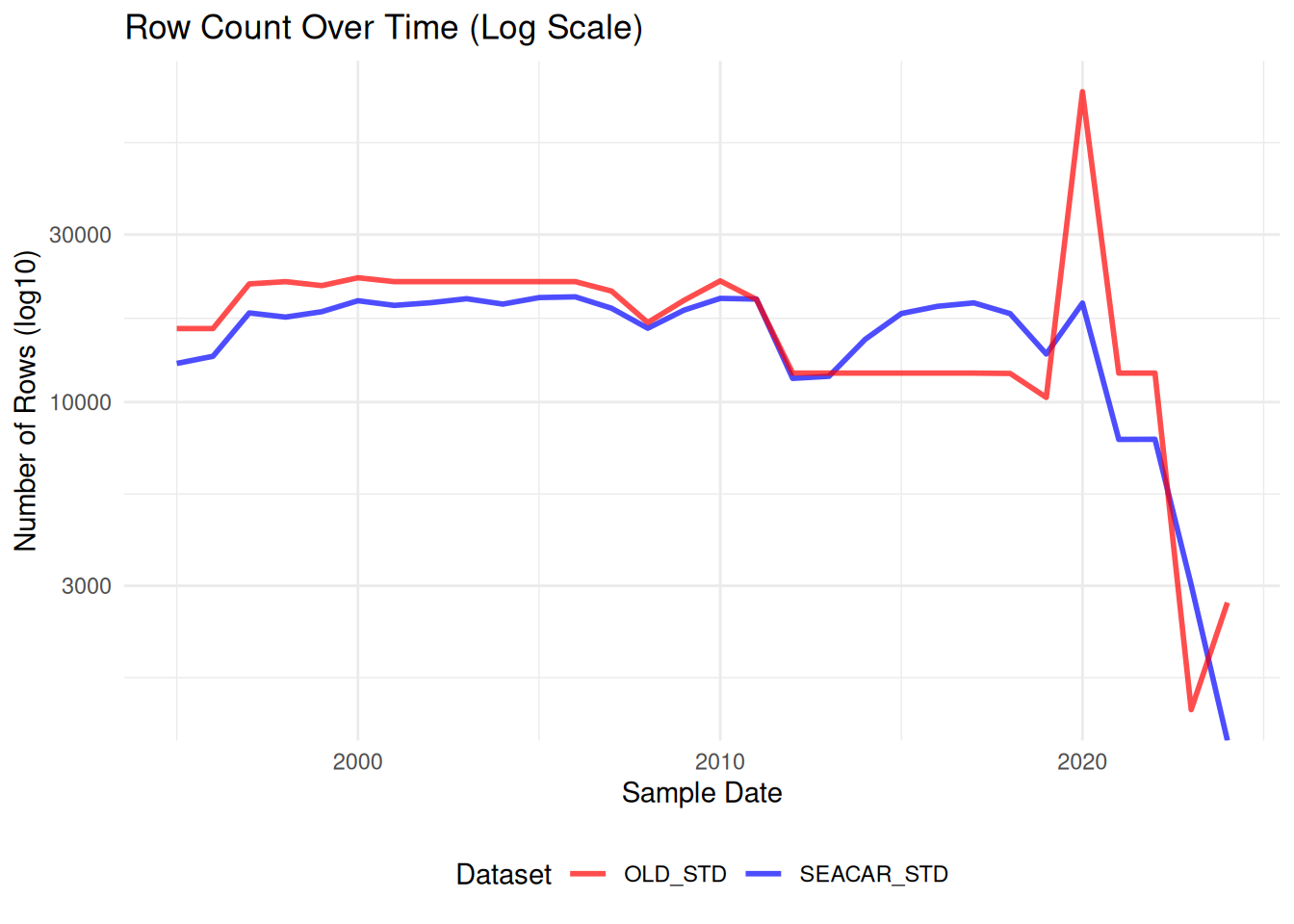
ggplot(combined_counts, aes(x = SampleDate)) +
geom_line(aes(y = SEACAR_Count, color = "SEACAR_STD", linetype = "SEACAR_STD"), size = 1, alpha = 0.7) +
geom_line(aes(y = OLD_Count, color = "OLD_STD", linetype = "OLD_STD"), size = 1, alpha = 0.7) +
scale_y_log10() +
scale_linetype_manual(values = c("SEACAR_STD" = "solid", "OLD_STD" = "solid")) +
scale_color_manual(values = c("SEACAR_STD" = "blue", "OLD_STD" = "red")) +
labs(
title = "Row Count Over Time (Log Scale)",
x = "Sample Date",
y = "Number of Rows (log10)",
color = "Dataset",
linetype = "Dataset"
) +
theme_minimal() +
theme(legend.position = "bottom")